I wrote this webapp to follow the dice roll distribution during a Catan game. Of course it can works with any game that uses from 1 to 3 dices.
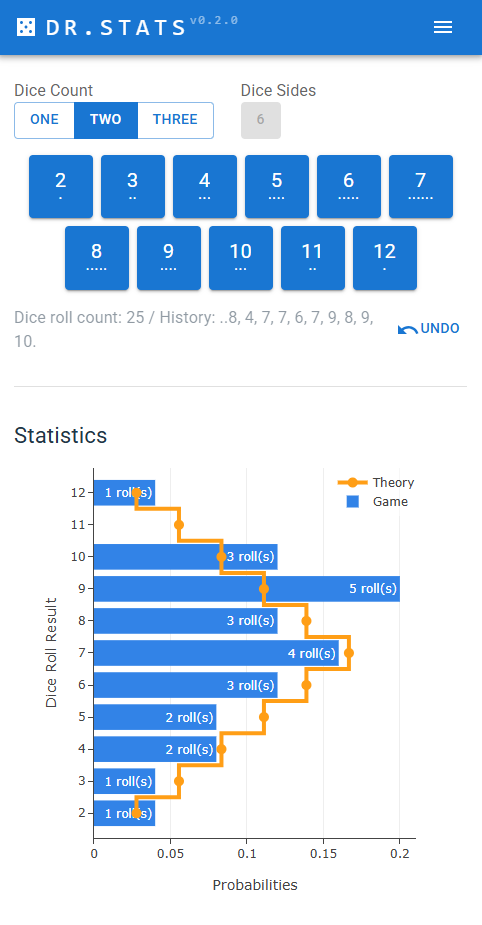
Each time dices are thrown, user can register it by pressing one of the blue buttons. This has for effect to update in real time the game statistics as blue bars. The theoretical values are the orange line.
User can visualize the history, and undo in case of errors. The state of the application is automatically saved/restored from the browser’s local storage.

React / Vite / Plotly.js / Material Design